Design of Experiments
I work a lot on Design of Experiments (DoE) though a lot of my work in this space is not public. I have been advising clients in the Gene Therapy and Pharma space on DoE. To get a sense of my DoE research on polymers, please see the paper linked below.
Machine learning in molecular engineering
My team innovated and publish on cutting-edge research in molecular engineering. Ongoing confidential projects relate to Latent Conditional Diffusion Models and protein/ RNA Foundation models.
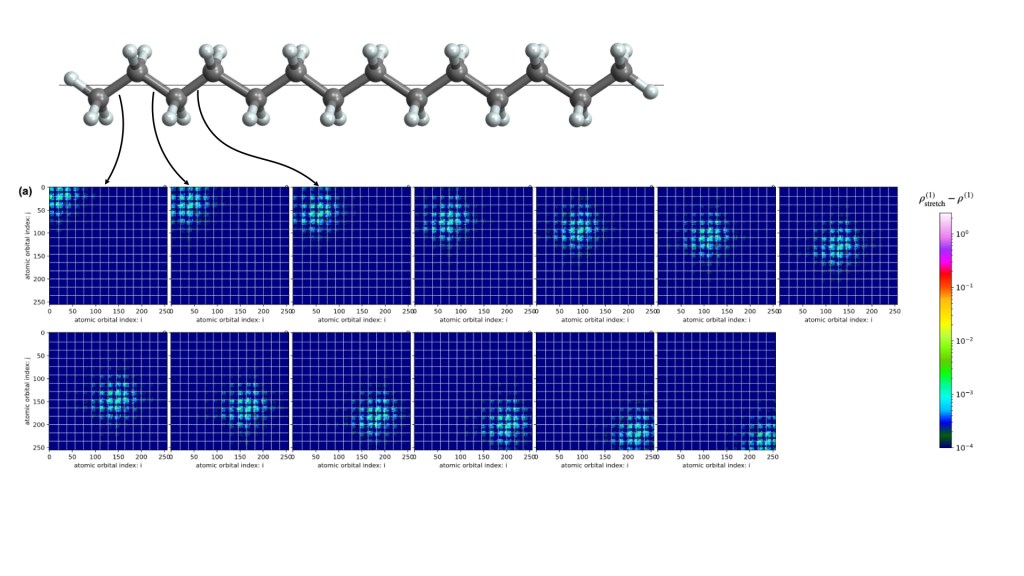
Past work relevant to molecular engineering: Machine Learning 1-RDM and 2-RDM.
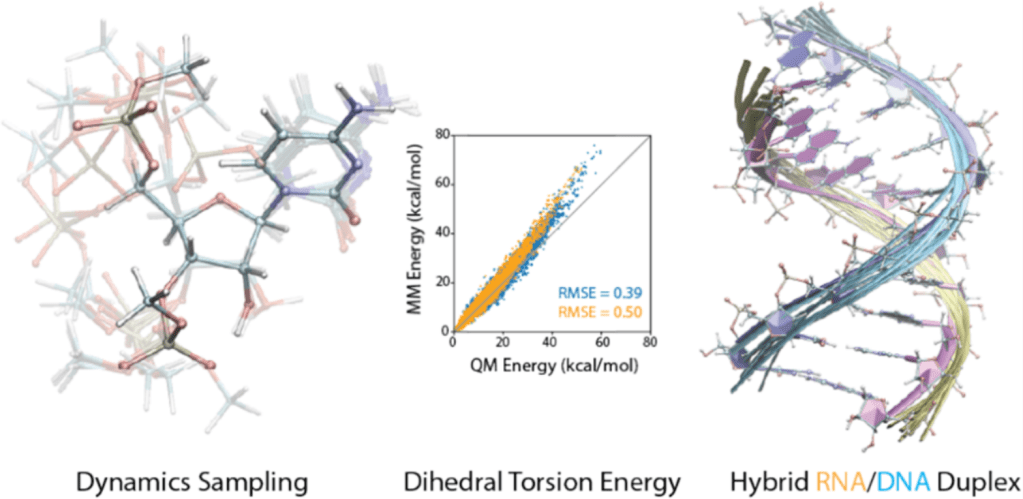
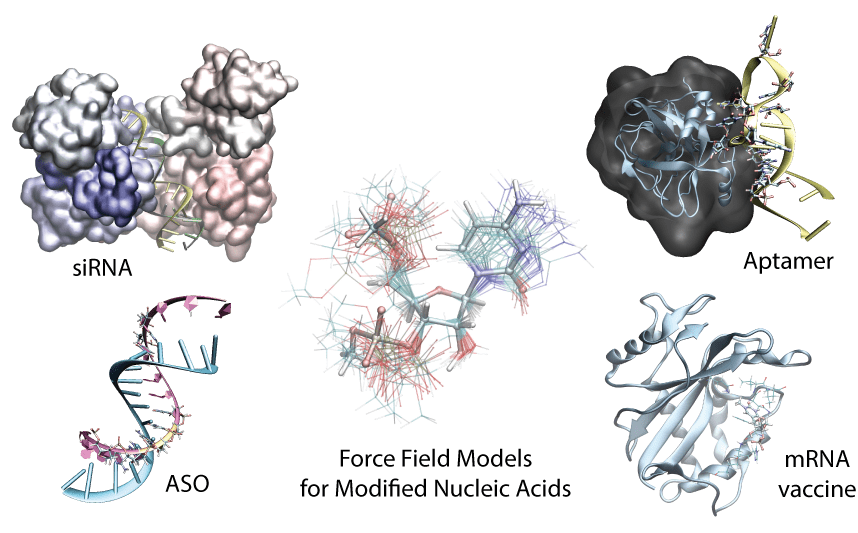
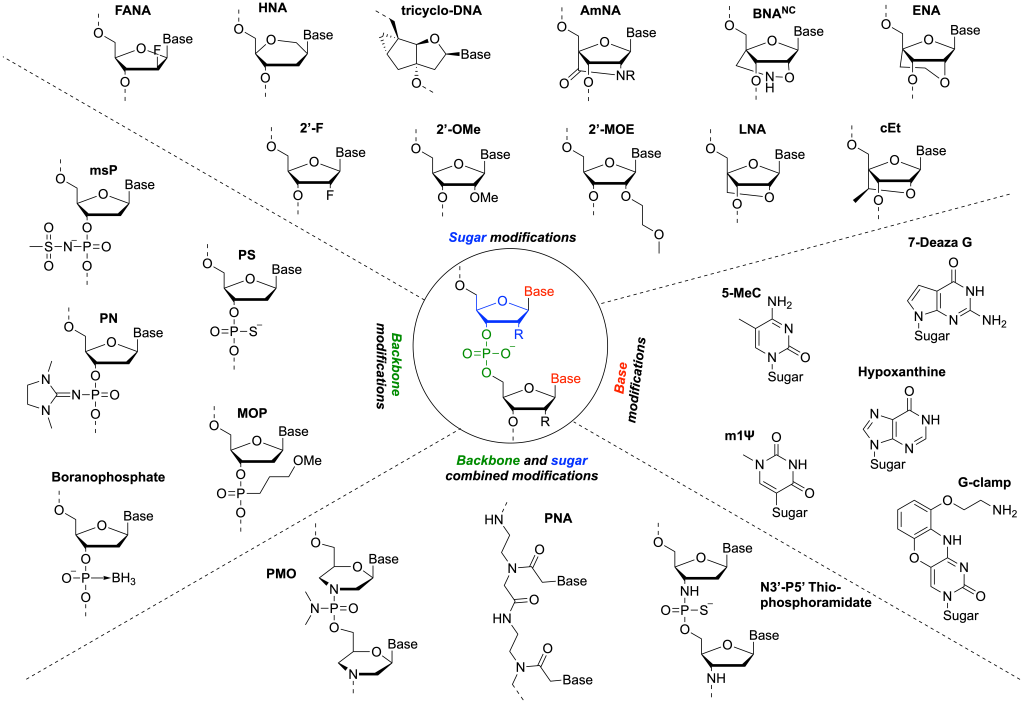
Generalized Methods to create Force Field Models
We introduced a unified, extensible framework for learning torsional energy landscapes in nucleic acids, pushing beyond traditional AMBER/CHARMM parameterization. The resulting force field delivers near-state-of-the-art accuracy while generalizing to chemically modified RNA/DNA systems.
NucleoPhi
We built NucleoPhi, a unified and transferable force-field framework for simulating chemically modified RNA and DNA at atomistic resolution, enabling simulation-driven design of therapeutic oligonucleotides.